Inhibition of growth and metastasis of triple-negative breast cancer targeted by Traditional Chinese Medicine Tubeimu in orthotopic mice models
Introduction
Breast cancer accounts for nearly one fourth of cancer cases worldwide and is the dominant cause of cancer death in females in China (1,2). Triple-negative breast cancer (TNBC) lacks the estrogen receptor (ER), progesterone receptor (PR), and human epidermal growth factor receptor 2 (HER-2) and accounts for 10%–20% of all breast cancer cases (1,3,4). The treatment for TNBC patient is very limited since it’s a metastatic recalcitrant cancer (5-7) with very poor prognosis (8-12). Therefore, transformative therapeutics are needed for TNBC.
Tubeimu (TBM), the rhizome of Bolbostemma paniculatum (Maxim.) Franquet, is a Traditional Chinese Medicinal (TCM) herb that has been used for the treatment of breast diseases, including breast cancer, for over 270 years (13). TBM has many pharmacological activities, including antitumor and anti-inflammatory efficacy (14,15), and has shown suppressive activity against several cancer cell lines, including nasopharyngeal (16) and liver (17,18) cancer cells.
The human breast cancer cell line MDA-MB-231 is derived from TNBC pleural effusion (19,20) and is invasive (21,22). In a previous study, we has demonstrated that the dichloromethane extract of TBM (ETBM) induced apoptosis of MDA-MB-231 TNBC cells in both two-dimensional and three-dimensional culture (23), but further studies are to be conducted to investigate the efficacy of ETBM on in vivo tumor formation and cell functions.
Therefore, we designed the present study to evaluate the efficacy of ETBM on different TNBC cell lines and MDA-MB-231 mice model. An orthotopic animal model of MDA-MB-231 was generated as it would enable a more clinical-like pattern of metastasis (24,25), while cancer cells expressing green fluorescent protein (GFP) enable real-time visualization and easier quantification of tumor growth (26). We also performed mRNA sequencing to have a more comprehensive understanding of which biological pathways ETBM might have influenced. Finally, quantitative real-time polymerase chain reactions (qRT-PCR), Western blotting and immunohistochemical (IHC) staining were used to confirm the change of expression on Rho GTPase activating protein 5 (ARHGAP5), integrin β1 (ITGβ1), and integrin β8 (ITGβ8) in focal adhesion pathways.
Materials and methods
Cell lines
MDA-MB-231-GFP human TNBC cells were obtained from AntiCancer, Inc. (San Diego, CA, USA) and cultured in RPMI-1640 (Gibco-BRL, Life Technologies, Inc., Carlsbad, CA, USA). MDA-MB-468 and SUM-149 were originally obtained from American Type Culture Collection (ATCC, Manassas, VA, USA) and cultured in Dulbecco’s modified eagle medium (DMEM)-high glucose (Gibco, Thermo Fisher Scientific, MA, USA). The 10% fetal bovine serum (FBS; Gibco) and penicillin-streptomycin antibiotic mixture (10 ml/L; Gibco) were added to the medium and cells were incubated in a 5% CO2 atmosphere at 37 °C.
ETBM agent
Fresh rhizomes of Bolbostemma paniculatum were collected in the Anguo City of Hebei Province in China (38.398 56°N; 115.348 43°E; altitude 28 m). The plant was identified and extracted with dichloromethane by the Pharmacy School of Beijing University of Chinese Medicine. Then it was dissolved in phosphate-buffered saline (PBS) with 10% of dimethyl sulfoxide (DMSO) for making a 1,000× ETBM solution. It was sterilized through a syringe with a 0.22 μm filter. ETBM was stored at –20 °C in eppendorf tubes and mixed with the medium for working concentrations. The 0.1% DMSO was used as control in in vitro studies.
Cell proliferation assay
In order to better visualize and quantitate ETBM activity on MDA-MB-231-GFP cells in vitro, we used the IncuCyte® ZOOM live-cell assay, which photographed each well on an hourly basis. Cells were plated into 96-well plates at 15,000 cells/well. Phase contrast and green fluorescence of the wells was monitored for a 24-h period. Concentrations of 1.875, 3.75, 7.5, 15, 30, 60, 120 and 480 μg/mL ETBM were applied on the cells. For half maximal inhibitory concentration (IC50) determination, MDA-MB-231, MDA-MB-468 and SUM-149 cells were seeded in 24-well plates and treated with increasing doses of ETBM in triplicate for 24 and 48 h. Then cells were stained with crystal violet. After washed and dried, stained cells were treated with lysate solution and shaken gently on a rocking shaker for 20–30 min. Diluted lysate solutions were added into 96-well plates and optical density (OD) was measured at 590 nm with BioTek ELx808 Absorbance Microplate Reader. IC50 values were calculated using nonlinear regression of log (ETBM’s concentration) vs. normalized inhibition rate.
Invasion/migration assay
Transwell inserts (Corning Inc., NY, USA) with 8 μm pore size with and without Matrigel (Collaborative Research Products, Bedford, MA, USA) were used. Cells were pretreated with 1% serum with no TBM, 30 μg/mL or 60 μg/mL TBM for 24 h and then mitomycin for 2 h. Following Matrigel rehydration with serum-free media, cells were seeded at a density of 104 per insert. The lower chamber was loaded with 450 μL RPMI-1640 containing 10% FBS. After 24 h incubation, cells that migrated through to the other side of the insert were fixed in 4% formalin, stained with 0.5% (weight/volume) crystal violet, and counted under a microscope.
Establishment of orthotopic breast cancer model
Animal experimental protocols were approved by the Animal Care and Protection Committee of Dongfang Hospital, Beijing University of Chinese Medicine (Approval No. 201540). First, five female nude mice (5–6 weeks old) were injected subcutaneously with a single dose of 5×106 MDA-MB-231-GFP human breast cancer cells. When the tumor size reached 1 cm3, the tumor tissue was harvested and cut into 1 mm3 fragments. Then, 40 female nude mice (5–6 weeks old) were implanted orthotopically with a single tumor fragment in the fat pad of the second left mammary gland.
Digital gene expression (DGE) sequencing
RNA extraction and preparation can be found in Supplementary Materials and Methods.
Clustering and sequencing (Novogene, Beijing, China): The clustering of the index-coded samples was performed on a cBot Cluster Generation System using a TruSeq PE Cluster Kit v3-cBot-HS (Illumina, San Diego, CA, USA) according to the manufacturer’s instructions. After cluster generation, the library preparations were sequenced on an Illumina Hiseq 2000/2500 platform, and 100 bp/50 bp single-end reads were generated.
Differential expression analysis (Novogene, Beijing, China): Differential expression analysis was performed using the DESeq R package (Version 1.10.1). DESeq provided statistical routines for determining differential expression in digital gene expression data using a model based on the negative binomial distribution. The resulting P-values were adjusted using the Benjamini and Hochberg’s approach for controlling the false discovery rate. Genes with an adjusted P<0.05 found by DESeq were assigned as differentially expressed. Then we applied pathway enrichment analysis using Kyoto Encyclopedia of Genes and Genomes (KEGG) with the Database for Annotation, Visualization and Integrated Discovery (DAVID). A heat map was drawn with Heml software (Version 1.0, Heatmap Illustrator, Wuhan, China).
qRT-PCR analysis
For mRNA detection, cDNAs were synthesized from 2 μg total RNA extracted from cells and tissues using oligo-d (T)15 primers and Moloney murine leukaemia virus (MMLV, Invitrogen, Thermo Fisher Scientific, MA, USA). qRT-PCR was performed by using SYBR Green PCR Master Mix (Toyobo, Osaka, Japan) on LightCycler® 480 Real-Time PCR System (Roche Molecular Diagnostics, Pleasanton, CA, USA). The primer sequences were as follows: glyceraldehyde-3-phosphate dehydrogenase (GAPDH) sense: 5’-GACCCCTTCATTGACCTCAAC-3’; GAPDH anti-sense: 5’-CTTCTCCATGGTGGTGAAGA-3’; ARHGAP5 sense: 5’-CATCTGTTTTTGGCCAACCT-3’; ARHGAP5 anti-sense: 5’-GTGGAGGAGCCACAATGTTT-3’; ITGB1 sense: 5’-AAATGGGCCTGTTCTTGATG-3’; and ITGB1 anti-sense: 5’-TTCTCCACAGTCCAGCAAGA-3’. The relative amount of genes was normalized to GAPDH. Data were calculated based on 2–ΔCt where ΔCt = CtTarget–CtReference. Fold change was calculated by the 2–ΔΔCt method.
Western blotting
Protein was extracted from cells and tissues using RIPA buffer containing a complete protease and phosphatase inhibitor cocktail (Roche, Mannheim, Germany). Protein (20 μg) was separated by electrophoresis on 8% sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) gels and blotted onto polyvinylidene fluoride (PVDF) membranes (Millipore, Darmstadt, Germany). Rabbit anti-ITGβ1 (1:2,000, Abcam, San Francisco, CA, USA); rabbit anti-ITGβ8 (1:1,000, Abcam); rabbit anti-ARHGAP5 (1:500, Abcam); rabbit anti-Phospho-Akt (Ser473) (1:1,000 dilution, Cell Signaling Technology, Danvers, MA, USA); on rabbit anti-Akt (1:1,000 dilution, Cell Signaling Technology), rabbit anti-GAPDH (1:1,000 dilution, Cell Signaling Technology) were used as primary antibodies. Horse radish peroxidase (HRP)-conjugated goat anti-rabbit (cwBiotech, Beijing, China) was used as the secondary antibody. Signals were visualized with chemiluminescence (Millipore, MA, USA).
IHC analysis
Frozen breast cancer tissues were embedded in optimal cutting temperature (OCT) compound (Sakura Finetek USA Inc., Torrance, CA, USA) and sliced using a cryostat. Slides were dried and fixed with 4% paraformaldehyde and stored in 4 °C. Slides were microwaved in 10 mM sodium citrate buffer before incubation with the primary antibodies against ARHGAP5 (1:300, Abcam) or ITGβ1 (1:400, Abcam) overnight at 4 °C. Slides were washed with PBS and incubated with streptavidin-horseradish peroxidase-conjugated goat secondary antibody (DAKO, Santa Clara, CA, USA) for 30 min at room temperature. The color was developed using diaminobenzidine. Finally, the slides were counter-stained with hematoxylin.
Statistical analysis
Each cell experiment was performed three times (n=3). The number of animals in each group was 10 (n=10). All experimental data were graphed and analyzed by GraphPad Prism 6 (GraphPad Software, Inc., San Diego, CA, USA). Comparisons between two groups were performed with two tailed unpaired t test. Welch correction was applied when necessary. Lung metastasis occurrence was analyzed with the χ2 test. Data are presented as x±s. P<0.05 were considered statistically significant.
More information on Material and Methods can be found in Supplementary Materials and Methods.
Results
ETBM inhibited TNBC cell proliferation in vitro
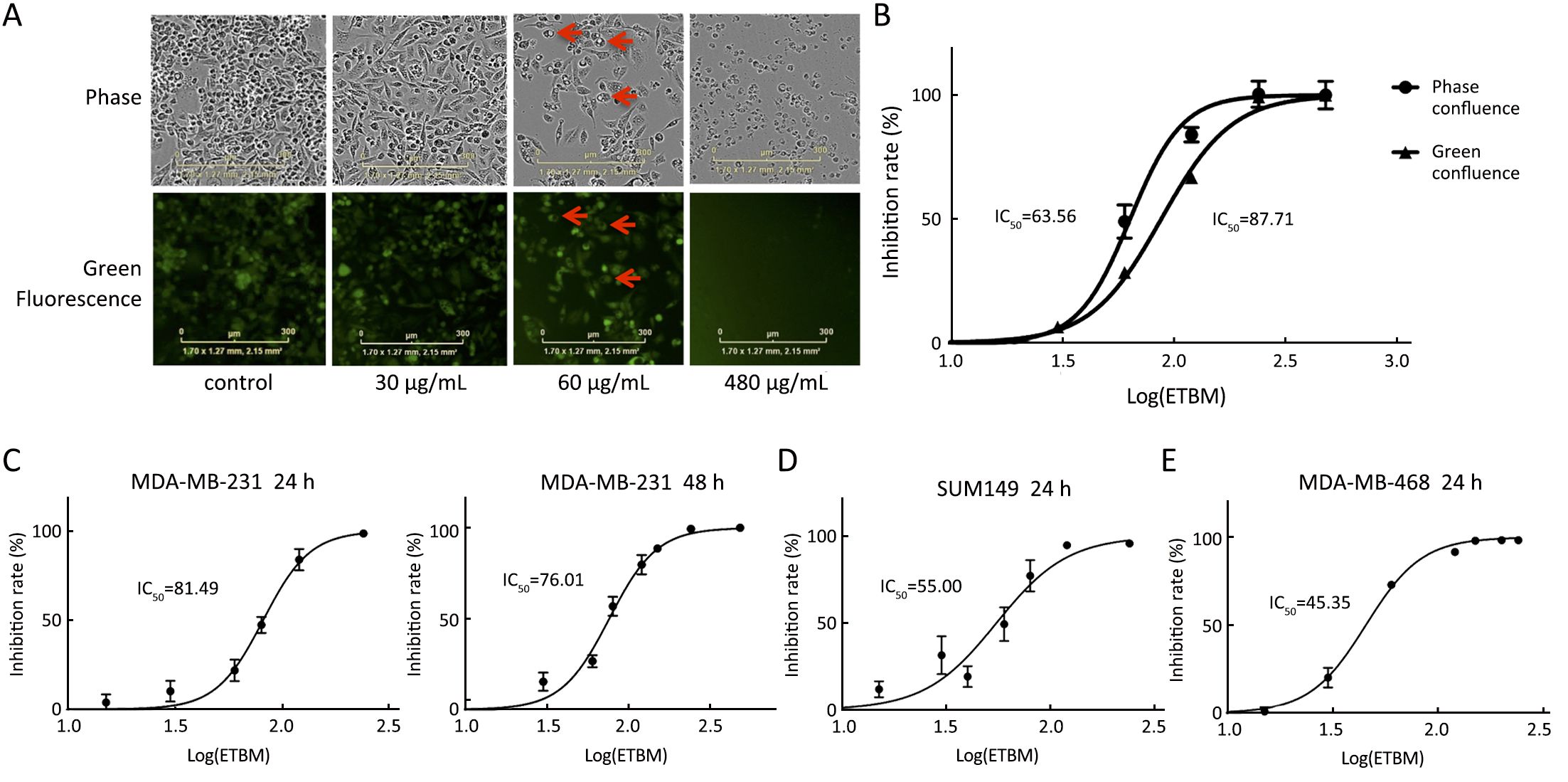
For better visualization on cell morphology, we adopted the IncuCyte® ZOOM live-cell assay. With pictures taken from IncuCyte® on 24 h of ETBM treatment, we could observe vigorously growing cells with GFP expression in the control wells, whereas only dead cells with disabled GFP expression in 480 μg/mL ETBM treatment wells (Figure 1A). Intriguingly, when treated with 60 μg/mL ETBM, cells generated many vesicles with no GFP expression (Figure 1A). We further calculated the IC50 on MDA-MB-231-GFP by IncuCyte®-based phase-confluence and fluorescence-confluence. And it was 63.56 μg/mL and 87.71 μg/mL, respectively (Figure 1B). Since the range is rather wide, we confirmed the IC50 adopting the crystal violet techniques. The estimated IC50 by crystal violet was 81.49 μg/mL on 24 h and 76.01 μg/mL on 48 h (Figure 1C). Furthermore, we investigated the IC50 of two different TNBC cell lines, SUM-149 and MDA-MB-468 (Figure 1D, E), which was 55.00 μg/mL and 45.35 μg/mL on 24 h, respectively. These results suggest that ETBM inhibits the proliferation of TNBC cell in a dose-dependent manner.
ETBM undermined the migration and invasion abilities of TNBC cells
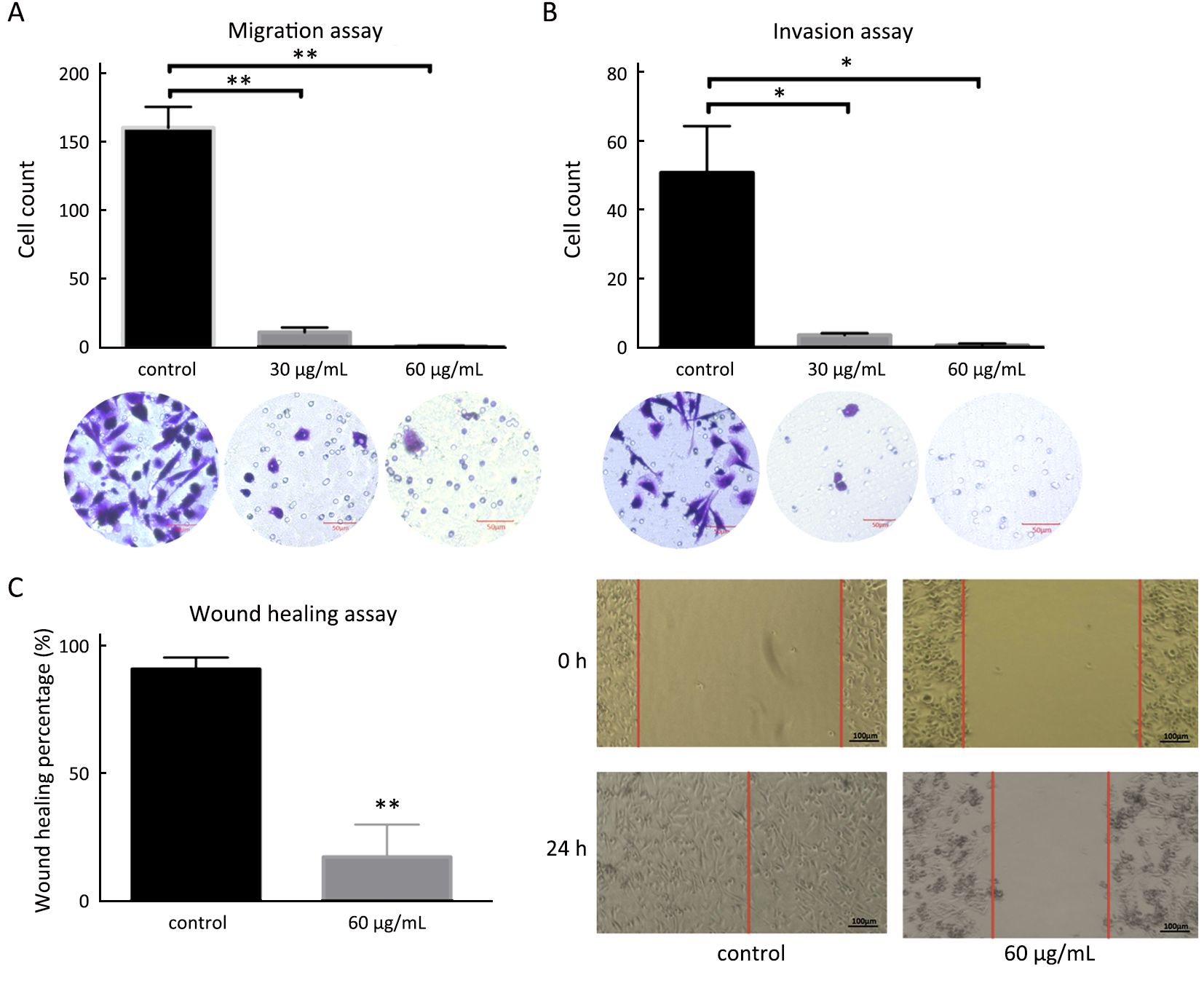
Migration and invasion of MDA-MB-231-GFP cells were inhibited by ETBM in 30 μg/mL and 60 μg/mL (Figure 2A, B). Migrating ETBM-treated cells became rounder with less focal adhesions than the control cells, where filopodia were often seen at the extremities. The wound-healing assay revealed that after 24 h, the wound in ETBM-treated cells healed much more slowly than that of untreated cells (P<0.01) (Figure 2C).
ETBM suppressed TNBC progression and lung metastasis in vivo
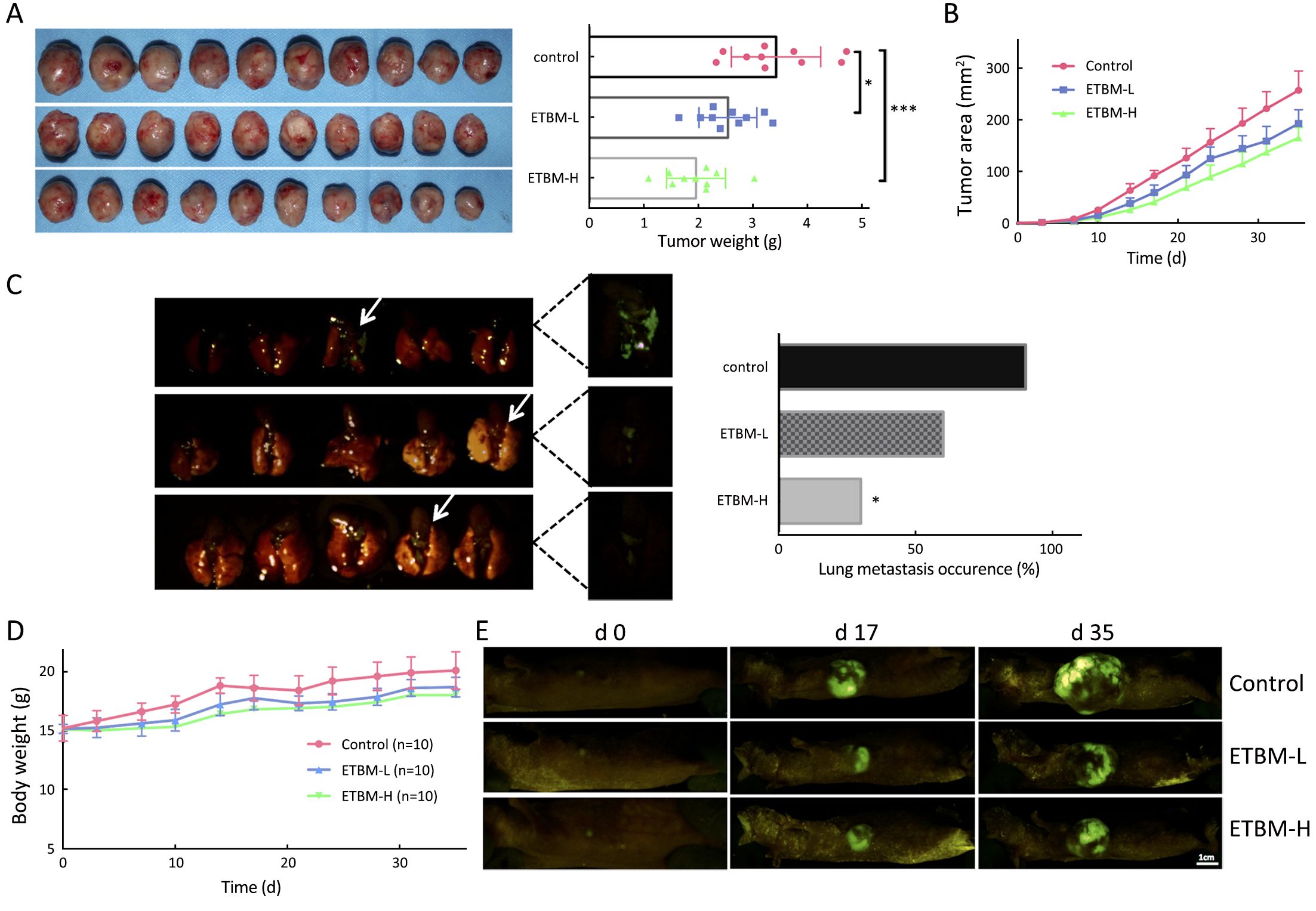
Tumors growth was inhibited in both high-dose (50 mg/kg) ETBM group (ETBM-H) and low-dose (30 mg/kg) ETBM group (ETBM-L) (Figure 3A, B, E). ETBM-H exhibited smaller tumor size compared to controls from d 10 (P<0.01). On sacrifice, tumors in ETBM-H displayed significantly lower weight compared to the untreated group (P<0.001). ETBM-L also showed meaningful inhibition on tumors (P<0.05). Moreover, ETBM-H notably suppressed lung metastasis occurrence (P<0.05) (Figure 3C).
Body weight and toxicity
During the course of the experiment, no physical or behavioral signs indicated adverse effects were observed in the animals in the ETBM treatment group. A stable body-weight gain from d 0 to d 35 in all groups was observed (Figure 3D).
Gene expression analysis of ETBM treated and untreated cells
Gene expression differences were detected in 665 genes, with 275 genes up-regulated and 390 genes down-regulated after ETBM treatment. KEGG analysis indicates that focal adhesion and extracellular matrix (ECM) might be significant pathways that were influenced by ETBM treatment (Figure 4A).
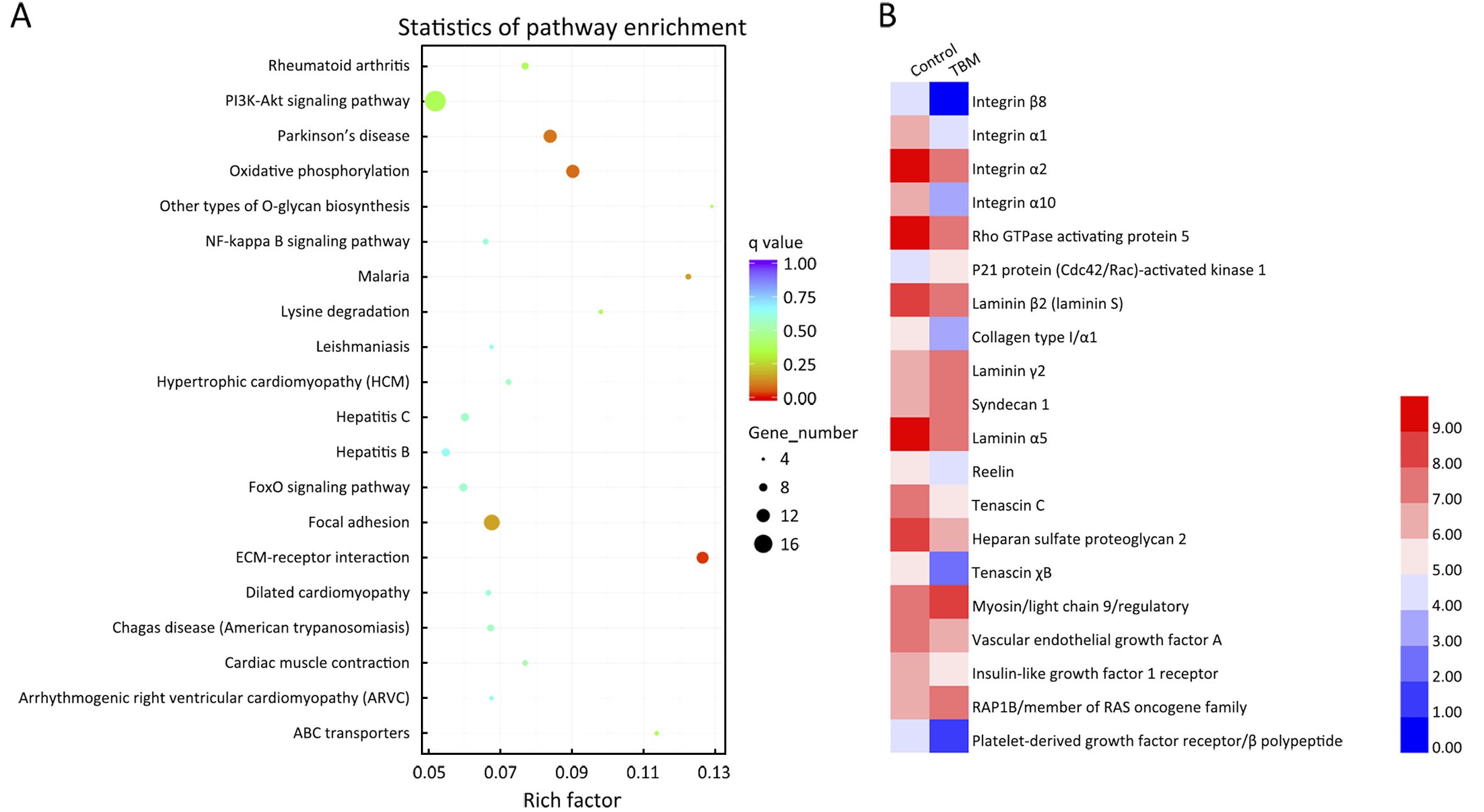
We then identified genes involved in these two pathways from the differential expression analysis and illustrated them on the heat map (Figure 4B). Integrins are the major transmembrane receptors mediating cell adhesion to ECM proteins (27) and ARHGAP5 (p190 RhoGAP) has an essential role in integrin-dependent adhesion signaling and cytoskeleton organization (28). Moreover, in TNBC, altered ECM can impair tumor’s response to therapeutics and promote progression along with metastasis (29). Therefore, we decided to investigate these proteins further to understand the biological mechanisms involved with ETBM treatment of TNBC.
Decreased expression of ARHGAP5 and integrins in ETBM-treated cell and tumors
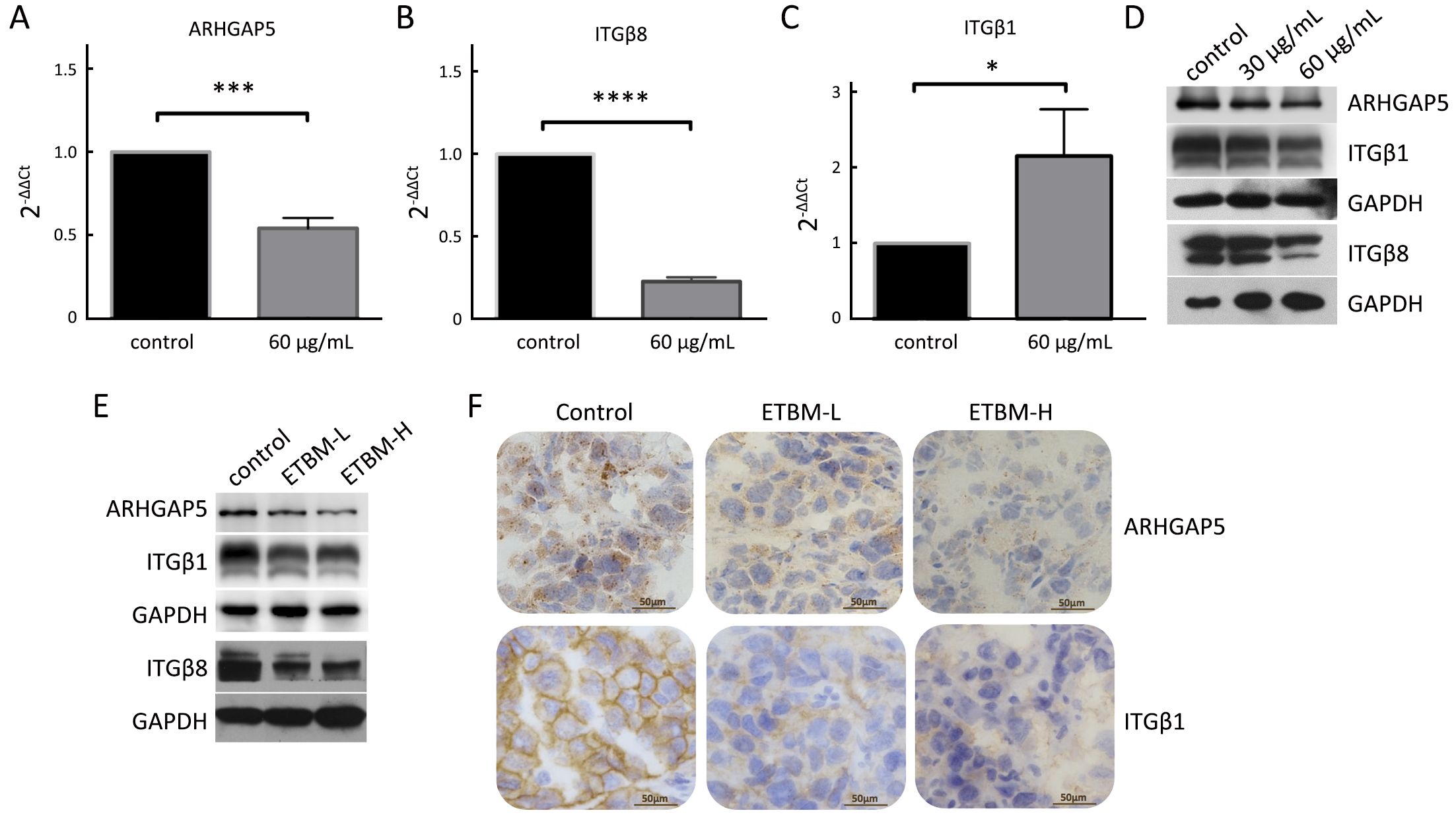
qRT-PCR and Western blotting were performed to further confirm the expression of ARHGAP5, ITGβ1 and ITGβ8 in ETBM treated cells (Figure 5A–D). There was reduction of mRNA and protein expression of ARHGAP5 and ITGβ8 in ETBM-treated cells, whereas ITGβ1 was decreased only at the protein level (Figure 5C, D). As for tumor samples, we observed suppressed expression of ARHGAP5, ITGβ1, and ITGβ8 in ETMB-treated groups (Figure 5E). Then IHC analysis verified reduction in ITGβ1 on the cell membrane and ARHGAP5 in the cytoplasm (Figure 5F). These data suggest that ETBM down-regulated these mobility-related proteins and this could contribute to the paralyzed migration and lesser metastases.
Discussion
According to Pharmacopoeia of the People’s Republic of China, the dosage of Tubeimu on human is 4.5 to 9.0 g. The main chemicals in Tubeimu are saponins, sterol compounds, alkaloids, maltol.I, e-modin.II, adenosine, etc. (30). In former studies regarding Tubeimu, one revealed that the aqueous extract from Tubeimu and Fuzi (Radix Aconiti Lateralis Preparata) had synergetic inhibitory effects on the growth of MDA-MB-231 cells and they obtained the same results with tubeimoside I and acetylbenzoylaconine, two well identified active compounds in these herbs (31). Our study adopted dichloromethane as extracting agent and fresh Tubeimu as samples for obtaining liposoluble chemicals, since our previous research showed the main compounds in ETBM are sterols instead of saponins (32). Besides the anti-proliferation effect, we observed inhibition on metastasis, cell migration and change in cell morphology, which may be caused by unidentified active compounds in ETBM. Furthermore, ETBM showed the efficacy without apparent toxicity. Low toxicity is among the advantages of using nature-developed medicine. ETBM has the potential to be developed into therapeutics for TNBC treatment. Developing an effective drug is a long-term endeavor and the exact active chemicals in ETBM need extensive study by multidisciplinary groups.
We illustrate that ETBM hindered the expression of ITGβ1, ITGβ8, and ARTHGAP5 in treated cells and tumors, which indicates that the decreasing cell motility and tumor metastasis may be a result of down-regulation of the integrin/ARHGAP5 pathway. ARHGAP5 protein was concentrated along filopodial shafts in cells, but was absent at the leading edge in lamellipodial protrusions (33). This could explain the morphological change we observed in the migrated cells. According to our study, the integrin/ARHGAP5 could be a potential target of ETBM. However, how the chemicals in ETBM interact with tumor ECM and influence the specific localization and quantity of integrins/ARHGAP5 needs further research.
By IncuCyte® photography, we observed plenty of intracellular vesicles in 60 μg/mL ETBM treated cells. Without access to electron microscope (EM), it was difficult to have a clear picture on what these vesicles contained. Nonetheless, in this case, they were unlikely to be transporting or secretory vesicles. We highly speculate that these vesicles were autophagosomes in the process of autophagy. The role of autophagy in cancer is controversial (34,35). It is necessary for cancer cells to adapt efficiently to hypo-nutrient environment, since protein degradation process leads to amino acid recycling and provides sufficient nutrients for cell survival (35). However, uncontrolled autophagy can induce autophagic cell death (ACD) (35). ACD is featured by the large-scale engulfment of portions of the cytoplasm in autophagosomes, giving the cell a special vacuolated appearance (36), like what we observed here. Autophagy can be destructive to cells by damaging the mitochondria, or degradation of vital proteins (37). The cells treated by ETBM lost viability and accumulated large vacuoles, which indicates autophagy may be among the mechanisms in the cause of their death. Autophagy related pathways, like phosphatidylinositol 3-kinase (PI3K)/Akt/mTOR, are to be studied in future studies.
Acknowledgements
We would like to thank Dr. Pan Deng at University of California, San Francisco (UCSF) for providing SUM-149 and MDA-MB-468 cell lines. The study was supported by National Natural Science Foundation of China Grant (No. 81303129) and Beijing University of Chinese Medicine Grant (Project ID: 2016-jxs-548).
Footnote
Conflicts of Interest: The authors have no conflicts of interest to declare.
Supplementary materials and methods
Wound healing assay
Cells were seeded at a density of 5×105 per well into a 24-well plate and allowed to reach confluence in culture. The cell monolayer was then scraped with a clean pipette tip to create an artificial wound. Images of the wound area were captured by photomicroscopy (Leica, Wetzlar, Germany) at 0 h and 24 h after injury.
Mice
Since MDA-MB-231 is derived from a female background, a total of 45 female BALB/C athymic nude mice (5–6 weeks old) were purchased from Beijing HFK Bioscience Co., Ltd. (Jing ICP 11027182). The animals were bred and maintained in a high efficiency particulate air (HEPA)-filtered environment with cages, food, and bedding sterilized by irradiation and autoclaving. All applicable international, national, and/or institutional guidelines for the care and use of animals were followed. All procedures performed in studies involving animals were in accordance with the ethical standards of the institution or practice at which the studies were conducted. Animal experimental protocols were approved by the Animal Care and Protection Committee of Dongfang Hospital, Beijing University of Chinese Medicine (Approval No.201540).
Dosing and schedule
On d 3 post-tumor implantation, the test agent was orally administered on a daily basis. The control group was given 0.1 mL 0.9% saline, while other groups were administered ETBM at doses of 30 mg/kg (low dose) and 50 mg/kg (high dose) of ETBM for 30 d consecutively.
Dosing and schedule
During the study, the animals were monitored for tumor growth and metastasis by whole-body fluorescence imaging using the FluorVivo Model 100 (INDEC BioSystems, Los Altos, CA, USA). Images of growing tumors were obtained twice a week. Mice were health checked daily. On d 35, all mice were sacrificed. The primary tumors were removed from each animal and weighed. Primary tumors and metastasis to the lungs were visualized by fluorescence imaging.
RNA extraction and preparation for sequencing
Total RNA from ETBM-treated and untreated cells (n=3) was extracted according to the protocol of the RNeasy Mini Kit (Qiagen, Valencia, CA, USA). RNA degradation and contamination were monitored on 1% agarose gels. RNA purity was determined using the NanoPhotometer® spectrophotometer (IMPLEN, Westlake Village, CA, USA). RNA concentration was measured using the Qubit® RNA Assay Kit in Qubit® 2.0 Flurometer (Life Technologies, Carlsbad, CA, USA). A total of 3 μg mRNA per sample was used as input material for the RNA sample preparations. Sequencing libraries were generated using the NEBNext® Ultra™ RNA Library Prep Kit for Illumina® (NEB, Ipswich, MA, USA) following the manufacturer’s recommendations. Index codes were added to attribute sequences to each sample. Briefly, mRNA was purified from total RNA using poly-T oligo-attached magnetic beads. Fragmentation was carried out using divalent cations under elevated temperature in NEBNext First Strand Synthesis Reaction Buffer (5X). First strand cDNA was synthesized using random hexamer primer and M-MuLV Reverse Transcriptase (RNase H-). Second strand cDNA synthesis was subsequently performed using DNA polymerase I and RNase H. Remaining overhangs were converted into blunt ends by exonuclease/polymerase. After adenylation of 3’ ends of DNA fragments, NEBNext Adaptor with a hairpin-loop structure was ligated to prepare for hybridization. In order to select cDNA fragments of preferentially 150–200 bp in length, the library fragments were purified with AMPure XP system (Beckman Coulter, Beverly, USA). USER enzyme (3 μL) (NEB) was then employed with size-selected, adaptor-ligated cDNA at 37 °C for 15 min followed by 5 min at
References
- Torre LA, Bray F, Siegel RL, et al. Global cancer statistics, 2012. CA Cancer J Clin 2015;65:87–108. [PubMed] DOI:10.3322/caac.21262
- Chen W, Zheng R, Zhang S, et al. Cancer incidence and mortality in China in 2013: an analysis based on urbanization level. Chin J Cancer Res 2017;29:1–10. [PubMed] DOI:10.21147/j.issn.1000-9604.2017.01.01
- Brenton JD, Carey LA, Ahmed AA, et al. Molecular classification and molecular forecasting of breast cancer: ready for clinical application? J Clin Oncol 2005;23:7350–60. [PubMed] DOI:10.1200/JCO.2005.03.3845
- Penault-Llorca F, Viale G. Pathological and molecular diagnosis of triple-negative breast cancer: a clinical perspective. Ann Oncol 2012;23 Suppl 6:vi19-22. [PubMed] DOI:10.1093/annonc/mds190
- Schneider BP, Winer EP, Foulkes WD, et al. Triple-negative breast cancer: risk factors to potential targets. Clin Cancer Res 2008;14:8010–8. [PubMed] DOI:10.1158/1078-0432.CCR-08-1208
- Yano S, Takehara K, Tazawa H, et al. Enhanced metastatic recurrence via lymphatic trafficking of a high-metastatic variant of human triple-negative breast cancer after surgical resection in orthotopic nude mouse models. J Cell Biochem 2017;118:559–69. [PubMed] DOI:10.1002/jcb.25735
- Li X, Yang J, Peng L, et al. Triple-negative breast cancer has worse overall survival and cause-specific survival than non-triple-negative breast cancer. Breast Cancer Res Treat 2017;161:279–87. [PubMed] DOI:10.1007/s10549-016-4059-6
- Cleator S, Heller W, Coombes RC. Triple-negative breast cancer: therapeutic options. Lancet Oncol 2007;8:235–44. [PubMed] DOI:10.1016/S1470-2045(07)70074-8
- Bauer KR, Brown M, Cress RD, et al. Descriptive analysis of estrogen receptor (ER)-negative, progesterone receptor (PR)-negative, and HER2-negative invasive breast cancer, the so-called triple-negative phenotype: a population-based study from the California cancer Registry. Cancer 2007;109:1721–8. [PubMed] DOI:10.1002/cncr.22618
- Reis-Filho JS, Tutt AN. Triple negative tumours: a critical review. Histopathology 2008;52:108–18. [PubMed] DOI:10.1111/j.1365-2559.2007.02889.x
- Criscitiello C, Azim HA Jr, Schouten PC, et al. Understanding the biology of triple-negative breast cancer. Ann Oncol 2012;23 Suppl 6:vi13-8. [PubMed] DOI:10.1093/annonc/mds188
- O’Shaughnessy J, Osborne C, Pippen JE, et al. Iniparib plus chemotherapy in metastatic triple-negative breast cancer. N Engl J Med 2011;364:205–14. [PubMed] DOI:10.1056/NEJMoa1011418
- Wang H. Wai Ke Zheng Zhi Quan Sheng Ji. Beijing: Chinese Medical Ancient Books Publishing House 1996;13:100.
- Zhao X. Ben Cao Gang Mu Shi Yi. Beijing: People’s Medical Publishing House, 1983:123-7.
- Yu TX, Ma RD, Yu LJ. Structure-activity relationship of tubeimosides in anti-inflammatory, antitumor, and antitumor-promoting effects.Acta Pharmacol Sin 2001;22:463–8. [PubMed]
- Weng XY, Ma RD, Yu LJ. Apoptosis of human nasopharyngeal carcinoma CNE-2Z cells induced by tubeimoside I. Ai Zheng (in Chinese) 2003;22:806–11. [PubMed]
- Wang Y, Deng L, Zhong H, et al. Natural plant extract tubeimoside I promotes apoptosis-mediated cell death in cultured human hepatoma (HepG2) cells. Biol Pharm Bull 2011;34:831–8. [PubMed] DOI:10.1248/bpb.34.831
- Yan ZC, Chen D, Wu XZ, et al. Effects of aqueous extracts of Aconitum carmichaeli, Rhizoma bolbostemmatis, Phytolacca acinosa, Panax notoginseng and Gekko swinhonis Gūenther on Bel-7402 cells. World J Gastroenterol 2007;13:2743–6. [PubMed] DOI:10.3748/wjg.v13.i19.2743
- Brinkley BR, Beall PT, Wible LJ, et al. Variations in cell form and cytoskeleton in human breast carcinoma cells in vitro. Cancer Res 1980;40:3118–29. [PubMed]
- Fernandez-Gallardo M, González-Ramírez R, Sandoval A, et al. Adenosine stimulate proliferation and migration in triple negative breast cancer cells. PLoS One 2016;11:e0167445. [PubMed] DOI:10.1371/journal.pone.0167445
- Schlienger S, Ramirez RA, Claing A. ARF1 regulates adhesion of MDA-MB-231 invasive breast cancer cells through formation of focal adhesions. Cell Signal 2015;27:403–15. [PubMed] DOI:10.1016/j.cellsig.2014.11.032
- Gest C, Joimel U, Huang L, et al. Rac3 induces a molecular pathway triggering breast cancer cell aggressiveness: differences in MDA-MB-231 and MCF-7 breast cancer cell lines. BMC Cancer 2013;13:63. [PubMed] DOI:10.1186/1471-2407-13-63
- Hu M, Zhao M, An C, et al. Real-time imaging of apoptosis induction of human breast cancer cells by the traditional Chinese medicinal herb tubeimu. Anticancer Res 2012;32:2509–14. [PubMed]
- Hoffman RM. Orthotopic metastatic mouse models for anticancer drug discovery and evaluation: a bridge to the clinic. Invest New Drugs 1999;17:343–59. [PubMed] DOI:10.1023/A:1006326203858
- Hoffman RM. Patient-derived orthotopic xenografts: better mimic of metastasis than subcutaneous xenografts. Nat Rev Cancer 2015;15:451–2. [PubMed] DOI:10.1038/nrc3972
- Hoffman RM. The multiple uses of fluorescent proteins to visualize cancer in vivo. Nat Rev Cancer 2005;5:796–806. [PubMed] DOI:10.1038/nrc1717
- Desgrosellier JS, Cheresh DA. Integrins in cancer: biological implications and therapeutic opportunities. Nat Rev Cancer 2010;10:9–22. [PubMed] DOI:10.1038/nrc2748
- Brouns MR, Matheson SF, Hu KQ, et al. The adhesion signaling molecule p190 RhoGAP is required for morphogenetic processes in neural development. Development 2000;127:4891–903. [PubMed]
- Yu T, Di G. Role of tumor microenvironment in triple-negative breast cancer and its prognostic significance. Chin J Cancer Res 2017;29:237–52. [PubMed] DOI:10.21147/j.issn.1000-9604.2017.03.10
- Sun J, Wen Q. Progress in medical studies of Bolbostemma paniculatum (Maxim.) Franquet. Zhongguo Yao Wu Jing Jie (in Chinese) 2010;7:430–1.
- Chen D, Cao R, He J, et al. Synergetic effects of aqueous extracts of Fuzi (Radix Aconiti Lateralis Preparata) and Tubeimu (Rhizoma Bolbostemmatis) on MDA-MB-231 and SKBR3 cells. J Tradit Chin Med 2016;36:113–24. [PubMed] DOI:10.1016/S0254-6272(16)30017-6
- Liu Y, Liu Y, Hu K, et al. In vitro study of fresh Tubeimu liposoluble extracts on breast cancer cells. Liao Ning Zhong Yi Za Zhi (in Chinese) 2013;40:768–9. DOI:10.13192/j.ljtcm.2013.04.166.liuyg.059
- Balanis N, Yoshigi M, Wendt MK, et al. β3 integrin-EGF receptor cross-talk activates p190RhoGAP in mouse mammary gland epithelial cells. Mol Biol Cell 2011;22:4288–301. [PubMed] DOI:10.1091/mbc.E10-08-0700
- Amaravadi R, Kimmelman AC, White E. Recent insights into the function of autophagy in cancer. Genes Dev 2016;30:1913–30. [PubMed] DOI:10.1101/gad.287524.116
- Yoshida GJ. Therapeutic strategies of drug repositioning targeting autophagy to induce cancer cell death: from pathophysiology to treatment. J Hematol Oncol 2017;10:67. [PubMed] DOI:10.1186/s13045-017-0436-9
- Kroemer G, Galluzzi L, Vandenabeele P, et al. Classification of cell death: recommendations of the Nomenclature Committee on Cell Death 2009. Cell Death Differ 2009;16:3–11. [PubMed] DOI:10.1038/cdd.2008.150
- Gozuacik D, Kimchi A. Autophagy and cell death. Curr Top Dev Biol 2007;78:217–45. [PubMed] DOI:10.1016/s0070-2153(06)78006-1
